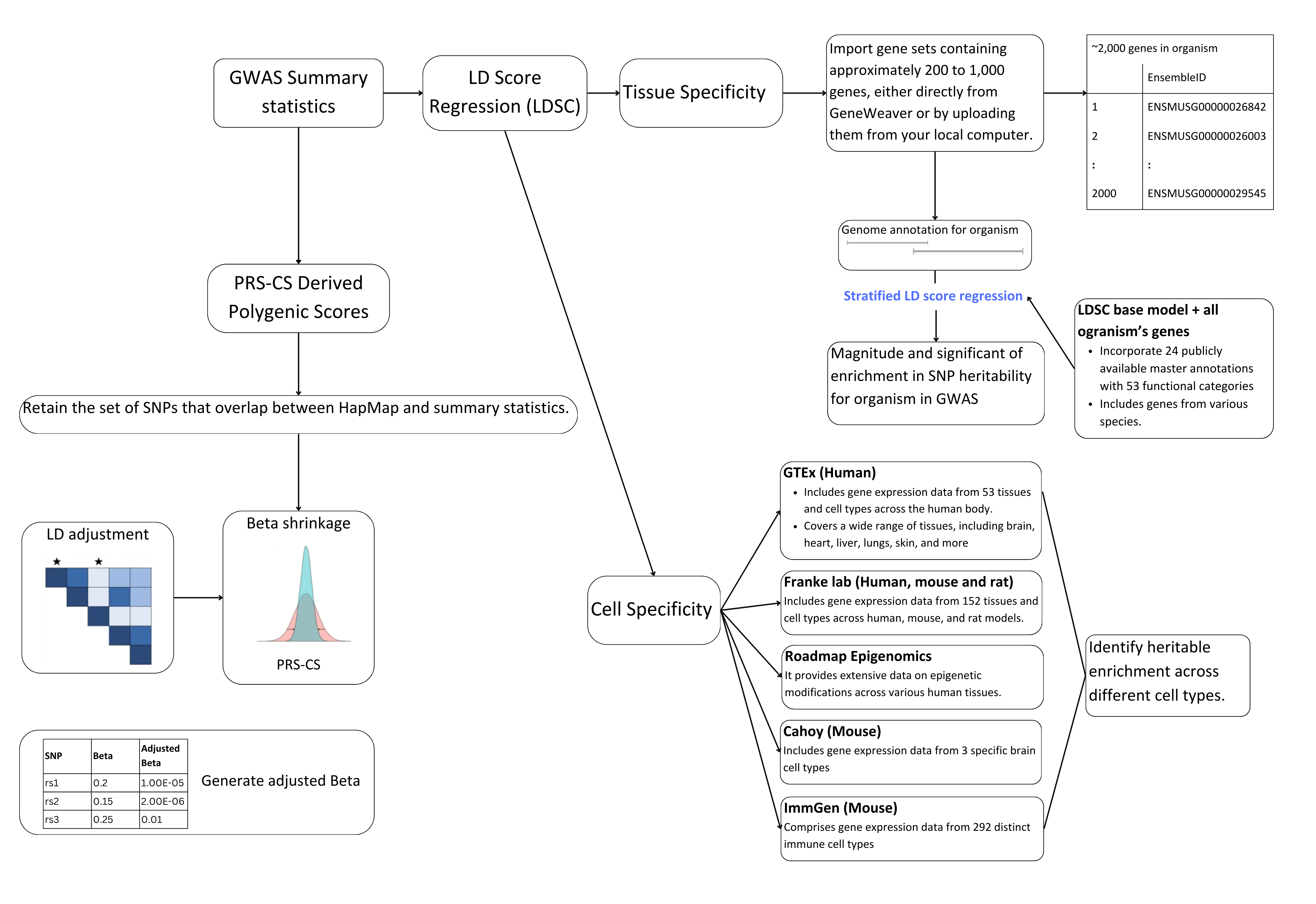
CONSILIENCE-GWAS: Web-based Enrichment Specificity and Polygenic Score Analysis enables users to investigate tissue/cell-specific enrichment and polygenic scores (PGS).
The platform calculates partitioned heritability, performs enrichment analysis for Genome-Wide Association Studies (GWAS),
and generates results highlighting the contribution of genetic variants across different functional categories.
Additionally, CONSILIENCE-GWAS allows cross-species enrichment analysis by integrating gene sets from model organisms (e.g., mouse, rat)
with human GWAS data, providing insights into how genetic signals from animal models relate to human traits.
Users receive detailed reports on tissue/cell-specific enrichments, cross-species comparisons, and the performance of PGS across various traits, with options to visualize and download the results.
Features
*CONSILIENCE-GWAS is free and open to all users and there is no login requirement.
*CONSILIENCE-GWAS was supported by a grant (DP1 DA042103) from the U.S. National Institutes of Health
Latest Updates
August 8, 2024
CONSILIENCE-GWAS 1.1 release (Integrated new GWAS summary statistics)
August 7, 2024
Bug fix (Resolved Trait Name error)
August 6, 2024
Bug fixes.
August 5, 2024
CONSILIENCE-GWAS 1.0 release (online enrichment analysis tool.)
1.Which build are the summary statistics and gene list based on?
Both the summary statistics and gene list are currently based on build 37.
We are in the process of updating to build 38.
2. Is the summary statistic based on the human or mouse genome?
The gene list is based on the mouse genome, while the summary statistics are based on the human genome.
3. How many megabytes can I upload?
You can upload files up to 100 megabytes in size.
Examples of Cross-species Functional Enrichment using LDSC
Huggett SB, Johnson EC, Hatoum AS, Lai D, Srijeyanthan J, Bubier JA, Chesler EJ, Agrawal A, Palmer AA, Edenberg HJ, Palmer RHC. Genes identified in rodent studies of alcohol intake are enriched for heritability of human substance use. Alcohol Clin Exp Res. 2021 Dec; 45(12):2485-2494. doi: 10.1111/acer.14738. Epub 2021 Dec 19. PubMed PMID: 34751961; PubMed Central PMCID: PMC9714312.
Benca-Bachman, Chelsie E., Jason Bubier, Rameez A. Syed, Pamela N. Romero Villela, and Rohan HC Palmer. "Polygenic influences on the behavioral effects of alcohol withdrawal in a mixed-ancestry population from the collaborative study on the genetics of alcoholism (COGA)." Molecular and Cellular Neuroscience 125 (2023): 103851.
Example of Cross-species Functional Enrichment using raw data via GCTA
Palmer RHC, Benca-Bachman CE, Huggett SB, Bubier JA, McGeary JE, Ramgiri N, Srijeyanthan J, Yang J, Visscher PM, Yang J, Knopik VS, Chesler EJ. Multi-omic and multi-species meta-analyses of nicotine consumption. Transl Psychiatry. 2021 Feb 4; 11(1):98. doi: 10.1038/s41398-021-01231-y. PubMed PMID: 33542196; PubMed Central PMCID: PMC7862377.
For Geneweaver:
For LDSC:
Bulik-Sullivan, Brendan K., et al. "LD Score regression distinguishes confounding from polygenicity in genome-wide association studies." Nature genetics 47.3 (2015): 291-295.
Finucane, Hilary K., et al. "Partitioning heritability by functional annotation using genome-wide association summary statistics." Nature genetics 47.11 (2015): 1228-1235.